※ GPS-Palm INTRODUCTION:
As an important and special class of post-translational modifications (PTMs), lipid modifications mainly comprise S-palmitoylation (C16), N-myristoylation (C14), S-farnesylation (C15), S-geranylgeranylation (C20), cholesterylation and glycosylphosphatidylinositol (GPI)-anchor, depending on the type of lipids covalently attached to modified substrate proteins (Ray, et al., 2017; Casey , 1995). Unlike other tethering lipid modifications, S-palmitoylation reversibly adds one or multiple palmitoyl moieties to internal cysteine residues in proteins through the thioesterification reaction (Roth, et al., 2006; Dietrich and Ungermann, 2004; Greaves and Chamberlain, 2007; Linder and Deschenes, 2007; Smotrys and Linder, 2004). S-palmitoylation effectively increases the hydrophobicity of protein surfaces to dynamically regulate membrane-protein interactions (Ray, et al., 2017; Kleuss and Krause , 2003) , and participates in regulating a broad spectrum of biological processes, such as signal transduction (Ray, et al., 2017; Smotrys and Linder, 2004), neuronal transmission (Roth, et al., 2006), metabolism (Shen, et al., 2017), autophagy (Kim , et al., 2019), and immunological response (Yao, et al., 2019). In addition, dysregulation of S-palmitoylation is associated with numerous human diseases such as cancer (Yao, et al., 2019; Chen, et al., 2017), neurodegenerative disorders (Andrew, et al., 2017) and diabetes (Berchtold, et al., 2011). Although many efforts have been made in this field, the experimental identification of S-palmitoylated proteins is tedious and laborious and the underlying molecular mechanisms are still unclear (Linder et al., 2007).
In this work, through the literature biocuration and public database integration, we compiled a large benchmark data set containing 3098 unique and non-homologous S-palmitoylation sites in 1682 proteins, which were experimentally identified from small- or large-scale studies, We developed a new method named data quality discrimination (DQD) to measure data quality weights (DQWs), and observed that small-scale sites had significantly higher DQWs than large-scale sites. We incorporated DQD into our recently developed GPS 5.0 algorithm, which implemented two additional methods of position weight determination (PWD) and scoring matrix optimization (SMO) for performance improvement, and achieved an area under the curve (AUC) value of 0.778 for predicting S-palmitoylation sites. Inspired by DeepVariant, a pioneering tool that encoded genomic sequencing data into images for calling genetic variants (Poplin, et al., 2018), we further designed a new strategy of number-to-image transformation (NIT) to transform numerical sequence similarity values into images, and such a method produced an increased AUC value of 0.806. We renamed this update of GPS algorithm as graphic presentation system 6.0. Additionally, we used NIT to encode six additional types of sequence-derived features including pseudo amino acid composition (PseAAC), composition of k-spaced amino acid pairs (CKSAAP), orthogonal binary coding (OBC), physicochemical properties in the Amino Acid index database (AAindex), autocorrelation functions (ACF) and position-specific scoring matrix (PSSM), and three types of structural features including accessible surface area (ASA), secondary structure (SS) and backbone torsion angles (BTA). A deep learning framework of parallel convolutional neural networks (pCNNs) was implemented for training and for integrating up to 2835 individual features, and then we developed a new tool called GPS-Palm. Through a comparison of other existing tools, GPS-Palm exhibited a >31.34% improvement of AUC value (0.855 vs. 0.651). Taken together, we anticipate GPS-Palm might be a helpful tool to analyze S-palmitoylation, and all approaches used in this study can be extended to predict other types of PTM sites. The local packages of GPS-Palm were implemented in Python and can be downloaded at http://gpspalm.biocuckoo.cn/download.php while the previous version is also provided.
GPS-Palm User Interface
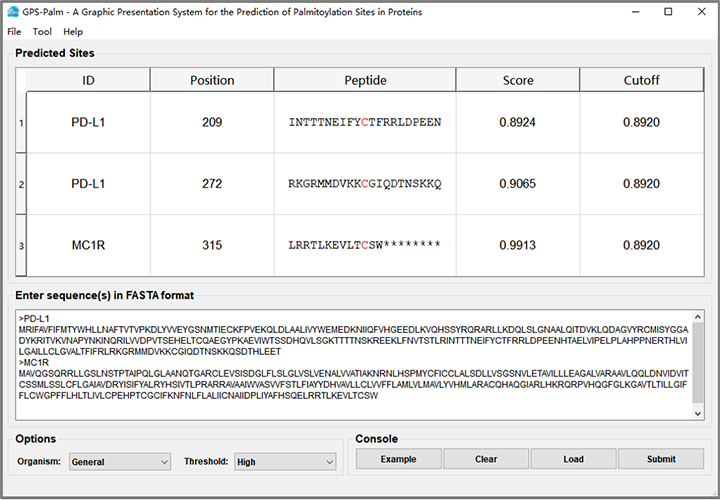
For publication of results please cite the following article:
|
|




 GPS-Palm: a deep learning-based graphic presentation system for the prediction of S-palmitoylation sites in proteins.
GPS-Palm: a deep learning-based graphic presentation system for the prediction of S-palmitoylation sites in proteins. CSS-Palm 2.0: an updated software for palmitoylation sites prediction.
CSS-Palm 2.0: an updated software for palmitoylation sites prediction.